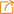A multi-omics integrative network map of maize
文献类型: 外文期刊
第一作者: Han, Linqian
作者: Han, Linqian;Zhong, Wanshun;Qian, Jia;Jin, Minliang;Tian, Peng;Zhu, Wanchao;Sun, Yonghao;Feng, Jia-Wu;Chen, Guo;Farid, Babar;Li, Ruonan;Xiong, Zimo;Li, Juan;Luo, Zi;Du, Dengxiang;Chen, Sijia;Jin, Qixiao;Li, Jiaxin;Li, Zhao;Liang, Yan;Jin, Xiaomeng;Peng, Yong;Zheng, Chang;Chen, Ling-Ling;Li, Qing;Yan, Jianbing;Yang, Fang;Li, Lin;Han, Linqian;Zhong, Wanshun;Qian, Jia;Jin, Minliang;Tian, Peng;Zhu, Wanchao;Sun, Yonghao;Li, Juan;Luo, Zi;Jin, Qixiao;Li, Zhao;Li, Qing;Yan, Jianbing;Yang, Fang;Li, Lin;Zhang, Hongwei;Liu, Xiangguo;Ye, Xinnan;Yin, Yuejia;Chen, Guo;Farid, Babar;Tian, Zhihui;Chen, Hong;Li, Weifu
作者机构:
期刊名称:NATURE GENETICS ( 影响因子:30.8; 五年影响因子:37.4 )
ISSN: 1061-4036
年卷期: 2023 年 55 卷 1 期
页码:
收录情况: SCI
摘要: Networks are powerful tools to uncover functional roles of genes in phenotypic variation at a system-wide scale. Here, we constructed a maize network map that contains the genomic, transcriptomic, translatomic and proteomic networks across maize development. This map comprises over 2.8 million edges in more than 1,400 functional subnetworks, demonstrating an extensive network divergence of duplicated genes. We applied this map to identify factors regulating flowering time and identified 2,651 genes enriched in eight subnetworks. We validated the functions of 20 genes, including 18 with previously unknown connections to flowering time in maize. Furthermore, we uncovered a flowering pathway involving histone modification. The multi-omics integrative network map illustrates the principles of how molecular networks connect different types of genes and potential pathways to map a genome-wide functional landscape in maize, which should be applicable in a wide range of species.
分类号:
- 相关文献
作者其他论文 更多>>
-
Acid-assisted ultrasonic preparation of nitrogen-doped MXene quantum dots for the efficient fluorescence "off-on-off" detection of Zn(II) in water and oxalic acid in vegetables
作者:Yang, Jinwen;Luo, Feili;Li, Lin;Cao, Feifei;Gu, Jiangjiang;Chen, Linlin;Qi, Jie;Wu, Honghong;Wu, Honghong;Wu, Honghong;Gu, Jiangjiang;Wu, Honghong;Gu, Jiangjiang
关键词:MXene quantum dots; Ultrasonic synthesis; Fluorescence sensors; Zn(II); Oxalic acid; Food samples
-
Lipids: A noteworthy role in better tea quality
作者:Huang, Fang-Fang;Bai, Si-Lei;Liu, Zhong-Hua;Li, Juan;Huang, Jian-An;Xiong, Li-Gui;Huang, Fang-Fang;Bai, Si-Lei;Liu, Zhong-Hua;Li, Juan;Huang, Jian-An;Xiong, Li-Gui;Bai, Si-Lei;Liu, Zhong-Hua;Li, Juan;Huang, Jian-An;Xiong, Li-Gui;Bai, Si-Lei;Liu, Zhong-Hua;Li, Juan;Huang, Jian-An;Xiong, Li-Gui;Yang, Pei-Di
关键词:Lipids; Preharvest tea leaves; Postharvest tea leaves; Flavor formation
-
Micronutrients and their effects on Horticultural crop quality, productivity and sustainability
作者:Ahmed, Nazir;Li, Juan;Xiao, Gengsheng;Wang, Qin;Hayat, Faisal;Tu, Panfeng;Zhang, Baige;Chachar, Zaid;Narejo, Mehar-un-Nisa;Bozdar, Bilqees;Deng, Lansheng
关键词:Nutrient deficiencies; Application methods; Yield enhancement; Fruit quality; Nutritional value; Postharvest preservation; Biofortification
-
Cryoprotective effect of fistular onion stalk polysaccharide on frozen surimi derived from bighead carp: Physicochemical properties and gel quality during storage
作者:Sun, Xiaoyue;Li, Qing;Ding, Ning;Liang, Shijie;Mubango, Elliot;Yu, Qinye;Dai, Ruitong;Tan, Yuqing;Luo, Yongkang;Hong, Hui;Zheng, Yanyan;Hong, Hui
关键词:Bighead carp surimi; Fistular onion stalk polysaccharide; Frozen storage; Cryoprotectant; Gel properties; Protein denaturation
-
Flavor precursors identification and thermal degradation mechanisms of glucoerucin in fragrant rapeseed oil
作者:Zhou, Qi;Zheng, Chang;Wei, Fang;Yang, Yini
关键词:Rapeseed; Widely targeted metabolism; Glucoerucin; Fragrant rapeseed oil; Thermally induced aroma; Models
-
Roles of substrates in removing antibiotics and antibiotic resistance genes in constructed wetlands: A review
作者:Cui, Erping;Gao, Feng;Cui, Erping;Zhou, Zhenchao;Chen, Hong;Li, Jianan
关键词:Comblned > Single soll; sand; gravel substrates; Others; speci& good removal; Constructed wetlands; Substrates; Antibiotics; Antibiotic resistance genes; Removal
-
A novel active transposon creates allelic variation through altered translation rate to influence protein abundance
作者:Chen, Guo;Wang, Ruilin;Jiang, Yizhe;Dong, Xiaoxiao;Xu, Jing;Xu, Qiang;Kan, Qiuxin;Luo, Zhixiang;Li, Qing;Chen, Guo;Springer, Nathan M.;Li, Qing
关键词: